ORIGINAL RESEARCH ARTICLE
Plasma and urine biomarker algorithm versus systematic biopsy for prostate cancer detection in elderly men: a randomised trial with early termination for futility
Torben Brøchner Pedersena  , Charlotte Aaberg Poulsenb
, Charlotte Aaberg Poulsenb  , Martin Lunda
, Martin Lunda  , Maher Albitarc
, Maher Albitarc  , Louise Dorner Østergaarda
, Louise Dorner Østergaarda  , Søren Feddersend
, Søren Feddersend  , Lars Lunda
, Lars Lunda  and Mads Hvid Poulsene,f
and Mads Hvid Poulsene,f 
aDepartment of Urology, Odense University Hospital, Odense, Denmark; bDepartment of Nuclear Medicine, Odense University Hospital, Odense, Denmark; cGenomic Testing Cooperative, Irvine, CA, USA; dDepartment of Clinical Biochemistry, Odense University Hospital, Odense, Denmark; eDepartment for Urology, Esbjerg Hospital, University Hospital of Southern Denmark, Esbjerg, Denmark; fDepartment of Regional Health Research, University of Southern Denmark, Esbjerg, Denmark
ABSTRACT
Objective: This study aimed to compare test sensitivity for detecting aggressive prostate cancer and test specificity (measured by reduction in prostate biopsies) between algorithm-triage and standard systematic biopsy in elderly men with suspected prostate cancer.
Methods: This randomised controlled trial enrolled men ≥ 70 years old suspected of prostate cancer and randomised them 1:1 to algorithm-triage or standard systematic biopsies. The algorithm arm used a 10-gene mRNA panel from urine and plasma samples, integrated with clinical characteristics and PSA measurements to predict prostate cancer with International Society of Urological Pathology grade group ≥ 2. Patient-reported outcomes measures were collected using the Functional Assessment of Cancer Therapy-Prostate (FACT-P) scores or subdomains throughout a 24-month follow-up. ClinicalTrials.gov: NCT04079699.
Results: A total of 202 men were included between October 2019 and September 2021. The study was terminated early due to algorithm underperformance. The algorithm-triage arm detected fewer indolent cancers (7.9% vs. 19%, absolute difference −10.9 percentage points, 95% confidence interval [CI]: −21.0 to −1.0 pp, P = 0.039) but also fewer clinically significant cancers (26% vs. 40%, absolute difference −13.9 percentage points, 95% CI: −27.6 to −0.1 pp, P = 0.051) compared to systematic biopsy. Patient-reported outcomes showed no significant between-group differences in FACT-P scores or subdomains throughout 24-month follow-up (differences 0.1–2.2 points, all P > 0.05), indicating comparable quality of life.
Conclusion: The biomarker-based algorithm-triage reduced biopsy numbers but also detected fewer clinically significant prostate cancers. Quality of life was comparable between approaches.
KEYWORDS: Prostate cancer; biomarkers; prostate biopsy
Citation: Scandinavian Journal of Urology 2026, VOL. 61, 31–37. https://doi.org/10.2340/sju.v61.45464.
Copyright: © 2026 The Author(s). Published by MJS Publishing on behalf of Acta Chirurgica Scandinavica. This is an Open Access article distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), allowing third parties to copy and redistribute the material in any medium or format and to remix, transform, and build upon the material, with the condition of proper attribution to the original work.
Received: 16 September 2025; Accepted: 13 January 2026; Published: 4 March 2026
CONTACT: Torben Brøchner Pedersen torben.b.pedersen@rsyd.dk Department of Urology, J. B. Winsløws Vej 4, Odense University Hospital, 5000 Odense, Denmark
Competing interests and funding: The authors have no conflicts of interest to disclose.
None.
Introduction
Prostate cancer remains a prevalent malignancy among men, necessitating accurate diagnostic methods for early detection and improved outcomes. Traditional diagnostic approaches including prostate-specific antigen (PSA) testing, digital rectal examination (DRE), and subsequent biopsy demonstrate notable limitations in specificity, potentially leading to overdiagnosis and overtreatment [1, 2]. The well-documented limitations of PSA in predicting aggressive disease have driven the development of novel biomarkers aimed at enhancing diagnostic accuracy [3]. Several novel biomarkers have emerged to improve diagnostic accuracy and reduce unnecessary biopsies. Urine-based biomarkers include PCA3 (Prostate Cancer Antigen 3), which offers superior specificity compared to PSA [4], and MyProstateScore (MPS) incorporating TMPRSS2-ERG gene fusion, demonstrating an area under the curve of 0.82 for detecting clinically significant prostate cancer [5]. SelectMDx and ExoDx Prostate Intelliscore have shown promising results while potentially reducing unnecessary biopsies [6, 7]. Blood-based biomarkers such as Prostate Health Index (PHI) and 4Kscore Test demonstrate superior performance compared to PSA alone, with studies showing 30–31% reduction in unnecessary biopsies while maintaining sensitivity for aggressive disease [8, 9]. Our research group developed a predictive algorithm utilising biomarkers from urine and plasma combined with clinical parameters. In a multicentre study, 524 men (median age 65 years; 77% Caucasian) who underwent prostate biopsy between 2014 and 2015 were evaluated; this cohort represented 21% of 2,528 initially enrolled men with prostate symptoms or elevated PSA. The algorithm was retrospectively applied to these biopsy results and was not used to determine whether biopsy was needed. It achieved 97% sensitivity and 97% negative predictive value for Gleason score ≥ 3+4, as well as 99% sensitivity and 99% negative predictive value for Gleason score ≥ 4+3 [10–13]. The approach could have reduced indolent cancer detection by 32% while missing only 0.8% of GS ≥ 7 cancers.
However, a parallel validation study among 314 biopsy-naïve men detected by unsystematic PSA screening (median age 68 years, median PSA 6.5 μg/L, enrolled between November 2018 and September 2021) revealed significantly lower performance, with only 87% sensitivity and 42% specificity for detecting ISUP GG ≥ 2 prostate cancer [14]. Clinical parameters alone outperformed the biomarker panel, highlighting challenges in translating promising algorithms to clinical practice.
Older men with suspected prostate cancer present a distinct clinical challenge. They are particularly vulnerable to procedural complications due to reduced physiological reserve and high comorbidity burden [15], and many may not benefit from aggressive diagnostic evaluation [16, 17]. Contemporary Nordic diagnostic guidance emphasises individualised approaches balancing detection of clinically significant disease against procedural burden [18]. We conducted a randomised controlled trial to evaluate whether a biomarker algorithm improves clinical decision-making in men aged 70 and older, enabling more individualised risk stratification while maintaining sensitivity for aggressive disease.
Patients and methods
The study was a randomised controlled single-centre trial of the algorithm and systematic biopsy in patients of age 70 years or above who were under suspicion of prostate cancer.
Inclusion criteria
Subjects were included in the study if they met all of the following criteria: suspicion of prostate cancer due to elevated PSA and/or palpable tumour in the prostate, indication for DRE and transrectal ultrasound with biopsy, age ≥ 70 years, PSA ≤ 20, and ability to provide informed written consent (competent adults only).
Exclusion criteria
Men were excluded from the study if they had been previously diagnosed with prostate cancer, were receiving treatment influencing PSA levels, had medical conditions that in the opinion of the investigator might interfere with the study (such as previous cancer-related therapy), had a life expectancy of less than 10 years, or had undergone prostate DRE within 24 h or ejaculation within 24 h prior to enrollment.
Randomisation (1:1) was used to assign patients to one of the following study arms: Arm A was the experimental arm where the algorithm determined whether to perform a prostate biopsy, and Arm B was the standard arm where every patient was offered a systematic prostate biopsy.
Subjects who fulfilled the inclusion criteria were asked to participate. An independent data manager from Odense Patient Explorative Network (OPEN) generated the permuted block allocation table, and the randomisation process was executed within the Research Electronic Data Capture (REDCap) database.
Study data were collected and managed using REDCap electronic data capture tools hosted at Odense University Hospital [19, 20].
Patients who were randomised to the standard arm followed the normal clinical evaluation with standard systematic prostate biopsy, 10 to 12 cores. Although multiparametric magnetic resonance imaging (mpMRI) was not standard of care for initial prostate cancer evaluation in biopsy-naïve men at study initiation, it was recommended by contemporary guidelines prior to repeat biopsy.
Patients who were randomised to the experimental arm had blood and urine samples collected and underwent transrectal ultrasound for determining prostate size. The biomarker analyses were then combined with the clinical data in the algorithm, as an initial diagnostic test. Based on the results, patients who were identified with a high-risk of having an aggressive cancer by the algorithm followed the normal procedures as described above, including prostate biopsies. The patients who were identified with a low-risk of having ISUP GG ≥ 2 prostate cancer according to the biomarker panel were not offered prostate biopsies; instead, they entered the follow-up directly. Secondary endpoint data were collected at 6, 12, and 24 months with Functional Assessment of Cancer Therapy-Prostate (FACT-P) questionnaire completion. Analysis and reporting were planned following completion of patient enrollment and the 2-year follow-up.
Urine and plasma processing
Each participant provided approximately 60 ml of first-void urine and 12 ml of peripheral whole blood (collected in two 6 ml EDTA vials) without undergoing a prior DRE. Urine was concentrated using Amicon Ultra-15 Centrifugal Filter Units (3 kDa cutoff) at 3220×g as previously described [11, 13]. Urine samples were collected in the morning from non-fasting participants, immediately placed on ice, and processed within 48 h.
Blood samples underwent centrifugation at 805 g for 10 min to separate plasma. Nucleic acid extraction for both RNA and DNA was performed using the NUCLISENS® easyMAG® Automated DNA/RNA extraction system (bioMérieux, Marcy-l’Étoile, France). After purification, nucleic acids were either immediately analysed or stored at −80°C for subsequent analysis. The RNA quality was maintained through strict adherence to protocols specifically designed to prevent degradation, with particular attention to storage and transport conditions.
Algorithm calculation
The biomarker test for estimating ISUP GG ≥ 2 prostate cancer probability has been comprehensively documented elsewhere [11, 12]. The test combines biomarker measurements with several clinical variables: patient age, prostate size, prior biopsy history, and PSA. These clinical factors are incorporated as independent variables within a logistic regression framework to enhance and calibrate the probability assessments for ISUP GG ≥ 2 prostate cancer. This integrated approach, combining biomarkers with clinical parameters through logistic regression, enables more accurate and patient-specific risk evaluations with greater clinical utility. While the test provides predictions at three sensitivity thresholds (high, medium, and low), our study utilised the high sensitivity threshold for ISUP GG ≥ 2 grade prostate cancer.
Primary endpoint
The two co-primary endpoints were: (1) test sensitivity, defined as the number of patients diagnosed with clinically significant prostate cancer (ISUP GG ≥ 2) in each study arm, and (2) test specificity, defined as the proportion of patients who underwent prostate biopsy in each arm. These endpoints assess whether the algorithm-triage approach maintains high sensitivity for detecting aggressive disease while improving specificity through reduction in unnecessary biopsies.
Secondary endpoints
Secondary endpoints included the number diagnosed with ISUP GG ≥ 3 prostate cancer and quality of life assessed using FACT-P at baseline, 6, 12, and 24 months post-randomisation. Subscale scores were considered valid when ≥ 50% of items were completed, with prorated scoring for missing items.
Sample size and statistical analysis
This randomised controlled trial was designed with two co-primary endpoints: (1) test sensitivity for detecting clinically significant prostate cancer (ISUP GG ≥ 2) and (2) test specificity measured by reduction in the proportion of patients undergoing prostate biopsy. Sample size calculation (α = 0.05, β = 0.2, 80% power) was based on the hypothesis of a 50% relative reduction in indolent prostate cancer detection (from 20% to 10%), which required 196 patients per arm [21, 22]. This hypothesis was informed by our previous multicentre study [10], which demonstrated that approximately 26% of the population would be classified as low-risk and potentially spared biopsy using the high-sensitivity threshold of the algorithm, while maintaining high sensitivity for clinically significant disease (ISUP GG ≥ 2).
We intended to enroll 350 patients per arm (700 total) to ensure adequate statistical power across more conservative effect size scenarios. A sensitivity analysis confirmed statistical power ≥ 95% with relative reductions as low as 40%.
We analysed the data using the intention-to-treat approach, with P < 0.05 considered statistically significant. Statistical analysis was performed using R version 4.5.2 (2025-10-31).
Analysis of detection and biopsy rates
To compare cancer detection and biopsy rates between the algorithm and systematic biopsy groups, we calculated the percentage of cases detected and percentage biopsied by each approach along with 95% confidence intervals. A two proportion z-test was used to calculate two-sided P-values and confidence intervals.
Analysis of patient-reported outcomes
Linear mixed-effects models were used to analyse longitudinal changes in FACT-P scores, with fixed effects for time point, treatment group, and their interaction, plus random subject intercepts. Models were fitted using restricted maximum likelihood estimation with diagnostic checks for normality and homoscedasticity. Estimated marginal means were calculated and compared between groups at each time point using Tukey’s method. A change of 6 points or more in FACT-P total score was considered clinically meaningful. Similar models were fitted for each subscale.
Patient comorbidity burden was assessed using a modified Charlson Comorbidity Index derived from diabetes, chronic obstructive pulmonary disease, ischaemic heart disease, dementia, other malignancy, and age-adjusted points, categorised as very low (0), low (1–2), moderate (3–4), or high risk (≥ 5).
Results
The study was terminated prior to reaching the target enrolment of 700 as initial monitoring suggested poor algorithm performance, which was later confirmed by our validation study [14]. Nevertheless, a total of 202 men were randomised between October 2019 and September 2021. Table 1 summarises the baseline demographics of both groups. Protocol deviations were documented as illustrated in the Figure 1. Three participants declined post-randomisation follow-up procedures but did not formally withdraw consent and were included in the intention-to-treat analysis.
| Variable | Algorithm | Standard biopsy | |
| Group size, n | 101 | 101 | |
| Age (IQR)/years | 75 (72, 77) | 76 (74, 78) | |
| Height (IQR)/cm | 176 (172, 180) | 178 (173, 182) | |
| Weight (IQR)/kg | 80 (74, 88) | 84 (75, 88) | |
| Body Mass Index (IQR)/kg/m2 | 26 (24, 28) | 26 (24, 28) | |
| Prostate volume (IQR)/mla | 54.0 (40.0, 72.0) | 58.5 (42.0, 82.5) | |
| PSA (IQR)/ng/ml | 8 (6, 12) | 8 (6, 12) | |
| PSA density (IQR)/ng/ml2 | 0.14 (0.10, 0.22) | 0.14 (0.09, 0.24) | |
| Patients with suspect DRE, n (%) | 18 (18%) | 32 (32%) | |
| Prebiopsy MRI, n (%) | 2 (2.0%) | 1 (1.0%) | |
| Previous negative biopsy, n (%) | 3 (3.0%) | 4 (4.0%) | |
| Charlson Risk Category Low | 1 (1.0%) | 0 (0.0%) | |
| Charlson Risk Category Moderate | 76 (75%) | 84 (83%) | |
| Charlson Risk Category High | 24 (24%) | 17 (17%) | |
| Data are presented as median with interquartile range (IQR) unless otherwise specified. IQR: interquartile range; DRE: digital rectal examination; MRI: magnetic resonance imaging; PSA: prostate-specific antigen. aProstate volume measured by transrectal ultrasound. |
|||
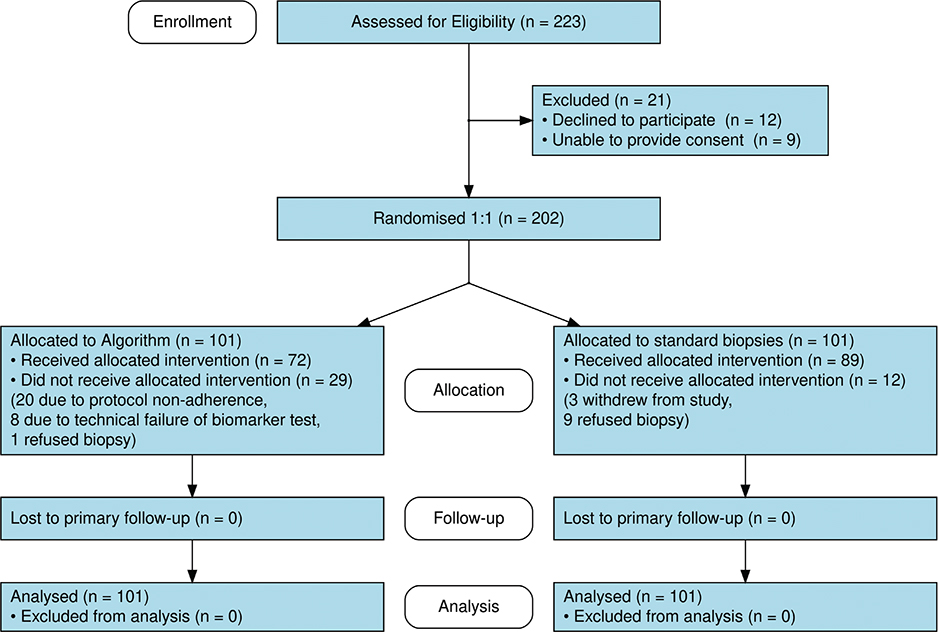
Figure 1. Consolidated Standards of Reporting Trials (CONSORT) flow diagram for a randomised controlled trial comparing biomarker algorithm-triage versus standard systematic prostate biopsy among men aged ≥ 70 years with suspected prostate cancer at Odense University Hospital, Denmark (October 2019–September 2021). The diagram illustrates enrollment (n = 223 assessed for eligibility), randomisation to algorithm-triage arm (n = 101) or systematic biopsy arm (n = 101), allocation and protocol adherence, follow-up completion, and analysis population for both arms.
Detection and biopsy rates
In the algorithm-triage arm, eight of 101 participants (7.9%) were diagnosed with indolent prostate cancer compared to 19 of 101 (19%) in the systematic biopsy group. The difference in detection rates between arms was −10.9 percentage points (pp) (95% confidence interval [CI]: −21.0 to −1.0 pp, P = 0.039). Regarding biopsy procedures, 57 of 101 participants (56%) in the algorithm group underwent prostate biopsies compared to 89 of 101 (88%) in the systematic biopsy arm. The difference in biopsy rates was −31.7 pp (95% CI: −44.2 to −19.1 pp, P < 0.001).
In the algorithm-triage arm, 26 of 101 participants (26%) were diagnosed with higher grade prostate cancer compared to 40 of 101 (40%) in the systematic biopsy group. The difference in detection rates between arms was −13.9 pp (95% CI: −27.6 to −0.1 pp, P = 0.051). A comprehensive breakdown of diagnostic outcomes stratified by algorithm prediction category is presented in Table 2.
For even higher grade disease (ISUP GG ≥ 3), the algorithm-triage arm identified 15 of 101 participants (15%) compared to 21 of 101 (21%) in the systematic biopsy group. The difference in detection rates for high-grade disease was −5.9 pp (95% CI: −17.0 to 6.0 pp, P = 0.358).
Table 3 summarises the distribution of algorithm results by biopsy status.
Only three men underwent prebiopsy mpMRI (one in the algorithm arm and two men in the systematic biopsy arm).
Quality of life outcomes
Linear mixed-effects analysis of FACT-P scores showed no statistically significant differences between treatment groups throughout the 24-month follow-up. The between-group difference in total FACT-P score at 24 months was 0.4 points (P = 0.225. All five FACT-P subdomains showed similarly minimal between-group differences (all P > 0.05).
Discussion
Our randomised controlled trial, while terminated prematurely, evaluated two co-primary endpoints: test sensitivity for detecting clinically significant prostate cancer and test specificity measured by reduction in unnecessary biopsies. Although the study achieved the specificity endpoint with a significant reduction in biopsy procedures (31.7 pp, P < 0.001), the algorithm failed to maintain non-inferior sensitivity for detecting aggressive prostate cancer. The algorithm-triage detected fewer clinically significant cancers (26% vs. 40%, P = 0.051) compared to standard biopsy, a finding that led to early termination and limits interpretation of the sensitivity endpoint.
However, the trial’s early termination due to the algorithm’s underperformance highlights a critical challenge in translational medicine: the gap between initial developmental promise and true clinical efficacy. While our algorithm showed exceptional sensitivity in earlier developmental studies [10], it failed to replicate this performance in a randomised, real-world setting, detecting fewer clinically aggressive cancers. This finding is corroborated by our recent validation study [14], which reported significantly lower performance metrics, with a negative predictive value of only 89%. This emphasises that a biomarker’s performance can be highly dependent on the patient population and clinical context in which it is applied.
The landscape of prostate cancer diagnostics has rapidly evolved, shifting towards integrated, risk-stratified pathways that incorporate imaging. Recent large-scale trials, such as the STHLM3-MRI study, have demonstrated that combining a risk prediction model with biparametric MRI and targeted biopsies can significantly reduce overdiagnosis while maintaining detection rates for clinically significant cancers [23–25]. Similarly, the PROMIS trial demonstrated the high negative predictive value of mpMRI, allowing for the avoidance of a substantial number of biopsies [26]. Recent advances include the development of 18-gene MyProstateScore version 2.0 (MPS2) showing 95% sensitivity and potential to avoid 40% of unnecessary biopsies [5], and the emergence of biparametric MRI approaches that demonstrate non-inferiority to multiparametric MRI while improving accessibility [27].
Contemporary developments in artificial intelligence offer additional promise. The MRI-based Predicted Transformer for Prostate Cancer (MRI-PTPCa) foundation model, trained on nearly 1.3 million image-pathology pairs, achieved area under the curve values above 0.978 with grading accuracy of 89.1% [28]. These findings suggest that a standalone biomarker algorithm, while useful for risk stratification, may be suboptimal as a primary biopsy decision-making tool without the inclusion of powerful imaging modalities such as MRI. Our results underscore this point, as the algorithm’s underperformance in a biopsy-naïve cohort suggests it may be better suited as a complementary tool within a more comprehensive diagnostic pathway.
No significant FACT-P differences were observed between groups, which is consistent with prior work [29]. Reducing biopsy numbers did not translate to measurable improvements in quality of life. However, FACT-P is designed for established cancer patients and may not capture diagnostic-specific distress in screening populations.
Our study has several important limitations. The premature termination resulted in a smaller sample size than originally planned, which reduced our statistical power, particularly for sub-analyses and less common outcomes such as aggressive disease. The single-centre design also limits the generalisability of our findings. However, a major strength is the study’s randomised design, which provides a high level of evidence for evaluating the clinical utility of a diagnostic algorithm, even in cases where the results are negative. The transparent reporting of algorithm failure is a critical scientific contribution, providing an important counterpoint to a field often dominated by positive findings from developmental studies.
Future research should focus on optimising diagnostic pathways by integrating multiple modalities. Combining refined biomarker algorithms with mpMRI in a sequential or parallel approach may offer the best balance of reducing unnecessary procedures while maintaining high sensitivity for higher grade disease. Larger, multicentre trials are needed to validate such integrated pathways across diverse patient populations.
In conclusion, the implementation of the biomarker-based decision algorithm resulted in fewer biopsies being taken, though this was accompanied with a lower detection of aggressive prostate cancers. When comparing quality of life as reported by patients, no substantial differences were observed between those who underwent algorithm-triage biopsies versus those who received standard systematic biopsies.
Acknowledgements
The authors gratefully acknowledge the Open Patient Data Explorative Network (OPEN) at Odense University Hospital, Denmark, for their data management support.
Claude Haiku 4.5 (Anthropic) was used to assist with language refinement and readability of the manuscript. Specifically, it was employed to improve sentence clarity and grammatical consistency throughout the manuscript. All authors critically reviewed and edited the AI-assisted content. The authors bear full responsibility for the accuracy, integrity, and scientific content of this work. AI technology was not used for literature review, generation of scientific insights, or formulation of conclusions.
Ethics statement
Approval was obtained from the Scientific Ethics Committee of the Region of Southern Denmark. Protocol Code number 69760.ClinicalTrials.gov Identifier: NCT04079699.
Authors’ contributions
Conception and design: MHP.
Data analysis and interpretation: TBP, MHP.
Data acquisition: MHP, LDØ.
Drafting the manuscript: TBP, MHP.
Critical revision of the manuscript for scientific and factual content: MHP, CAP, LL, MA.
Statistical analysis: TBP.
Supervision: MHP.
Data availability
The dataset generated and analysed during the current study is not publicly available, but are available from the corresponding author on reasonable request.
References
[1] Vickers A, O’Brien F, Montorsi F, et al. Current policies on early detection of prostate cancer create overdiagnosis and inequity with minimal benefit. BMJ. 2023;381:e071082. https://doi.org/10.1136/bmj-2022-071082
[2] Loeb S, Bjurlin MA, Nicholson J, et al. Overdiagnosis and overtreatment of prostate cancer. Eur Urol. 2014;65:1046–1055. https://doi.org/10.1016/j.eururo.2013.12.062
[3] David MK, Leslie SW. Prostate-specific antigen. StatPearls [Internet]. Treasure Island, FL: StatPearls Publishing; 2025 [cited 2025 Apr 22]. Available from: http://www.ncbi.nlm.nih.gov/books/NBK557495/
[4] Ploussard G, de la Taille A. The role of prostate cancer antigen 3 (PCA3) in prostate cancer detection. Expert Rev Anticancer Ther. 2018;18: 1013–1020. https://doi.org/10.1080/14737140.2018.1502086
[5] Tosoian JJ, Zhang Y, Xiao L, et al. Development and validation of an 18-gene urine test for high-grade prostate cancer. JAMA Oncol. 2024;10:726–736. https://doi.org/10.1001/jamaoncol.2024.0455
[6] Hendriks RJ, van der Leest MMG, Israël B, et al. Clinical use of the SelectMDx urinary-biomarker test with or without mpMRI in prostate cancer diagnosis: a prospective, multicenter study in biopsy-naïve men. Prostate Cancer Prostatic Dis. 2021;24:1110–1119. https://doi.org/10.1038/s41391-021-00367-8
[7] Margolis E, Brown G, Partin A, et al. Predicting high-grade prostate cancer at initial biopsy: clinical performance of the ExoDx (EPI) Prostate Intelliscore test in three independent prospective studies. Prostate Cancer Prostatic Dis. 2022;25:296–301. https://doi.org/10.1038/s41391-021-00456-8
[8] Nordström T, Vickers A, Assel M, et al. Comparison between the four-kallikrein panel and prostate health index for predicting prostate cancer. Eur Urol. 2015;68:139–146. https://doi.org/10.1016/j.eururo.2014.08.010
[9] Wagaskar VG, Sobotka S, Ratnani P, et al. A 4K score/MRI-based nomogram for predicting prostate cancer, clinically significant prostate cancer, and unfavorable prostate cancer. Cancer Rep (Hoboken). 2021;4:e1357. https://doi.org/10.1002/cnr2.1357
[10] Albitar M, Ma W, Lund L, et al. A multi-center prospective study to validate an algorithm using urine and plasma biomarkers for predicting Gleason ≥3+4 prostate cancer on biopsy. J Cancer. 2017;8:2554–2560. https://doi.org/10.7150/jca.20031
[11] Albitar M, Ma W, Lund L, et al. Predicting prostate biopsy results using a panel of plasma and urine biomarkers combined in a scoring system. J Cancer. 2016;7:297–303. https://doi.org/10.7150/jca.12771
[12] Ma W, Diep K, Fritsche HA, et al. Diagnostic and prognostic scoring system for prostate cancer using urine and plasma biomarkers. Genet Test Mol Biomarkers. 2014;18:156–163. https://doi.org/10.1089/gtmb.2013.0424
[13] Poulsen MH, Feddersen S, Albitar M, et al. A prospective study of a urine and plasma biomarker test for the prediction of gleason ≥ 3 + 4 prostate cancer in a mixed cohort. Scand J Urol. 2020;54:323–327. https://doi.org/10.1080/21681805.2020.1782979
[14] Pedersen TB, Poulsen MH, Lund M, et al. Prospective validation study of a combined urine and plasma test for predicting high-grade prostate cancer in biopsy naïve men. Scand J Urol. 2025;60:8–13. https://doi.org/10.2340/sju.v60.42752
[15] Brewster DH, Fischbacher CM, Nolan J, et al. Risk of hospitalization and death following prostate biopsy in Scotland. Public Health. 2017;142:102–110. https://doi.org/10.1016/j.puhe.2016.10.006
[16] Albertsen PC, Hanley JA, Fine J. 20-year outcomes following conservative management of clinically localized prostate cancer. JAMA. 2005;293:2095–2101. https://doi.org/10.1001/jama.293.17.2095
[17] Thomsen FF, Garmo H, Egevad L, et al. Temporal trend in risk of prostate cancer death in men with favourable-risk prostate cancer. Scand J Urol. 2024;59:76–83. https://doi.org/10.2340/sju.v59.34015
[18] Stranne J. 2023/2024 update of the national prostate cancer guidelines in Sweden. Scand J Urol. 2024;59:210–211. https://doi.org/10.2340/sju.v59.42656
[19] Harris PA, Taylor R, Minor BL, et al. The REDCap consortium: building an international community of software platform partners. J Biomed Inform. 2019;95:103208. https://doi.org/10.1016/j.jbi.2019.103208
[20] Harris PA, Taylor R, Thielke R, et al. Research electronic data capture (REDCap) – a metadata-driven methodology and workflow process for providing translational research informatics support. J Biomed Inform. 2009;42:377–381. https://doi.org/10.1016/j.jbi.2008.08.010
[21] Pocock SJ. Clinical trials with multiple outcomes: a statistical perspective on their design, analysis, and interpretation. Control Clin Trials. 1997;18:530–545; discussion 546–549. https://doi.org/10.1016/S0197-2456(97)00008-1
[22] Machin D, Campbell MJ, Tan S-B. Sample size tables for clinical studies. 3rd ed. Chichester, UK: Wiley; 2009.
[23] Lazarovich A, Vigneswaran H, Palsdottir T, et al. A comparison of Stockholm3, serum biomarkers, and risk calculators to predict prostate cancer in a racially and ethnically diverse cohort: evaluation of the Stockholm3 multiethnic SEPTA Trial. J Urol. 2025;213:590–599. https://doi.org/10.1097/JU.0000000000004437
[24] Nordström T, Discacciati A, Bergman M, et al. Prostate cancer screening using a combination of risk-prediction, MRI, and targeted prostate biopsies (STHLM3-MRI): a prospective, population-based, randomised, open-label, non-inferiority trial. Lancet Oncol [Internet]. 2021;22:1240–1249. https://doi.org/10.1016/S1470-2045(21)00348-X
[25] Vigneswaran HT, Eklund M, Discacciati A, et al. Stockholm3 in a multiethnic cohort for Prostate Cancer Detection (SEPTA): a prospective multicentered trial. J Clin Oncol. 2024;42:3806–3816. https://doi.org/10.1200/JCO.24.00152
[26] Ahmed HU, El-Shater Bosaily A, Brown LC, et al. Diagnostic accuracy of multi-parametric MRI and TRUS biopsy in prostate cancer (PROMIS): a paired validating confirmatory study. Lancet. 2017;389:815–822. https://doi.org/10.1016/S0140-6736(16)32401-1
[27] Ng ABCD, Asif A, Agarwal R, et al. Biparametric vs multiparametric MRI for prostate cancer diagnosis: the PRIME diagnostic clinical trial. JAMA [Internet]. 2025;334:1170–1179. https://doi.org/10.1001/jama.2025.13722
[28] Shao L, Liang C, Yan Y, et al. An MRI-pathology foundation model for noninvasive diagnosis and grading of prostate cancer. Nat Cancer. 2025;6:1621–1637. https://doi.org/10.1038/s43018-025-01041-x
[29] Shankar PR, Ellimoottil C, George AK, et al. Testing-related health impact of transrectal and transperineal prostate biopsy as assessed by health utilities. J Urol. 2021;206:1403–1410. https://doi.org/10.1097/JU.0000000000002118