ORIGINAL REPORT
Identifying Mild-to-Moderate Atopic Dermatitis Using a Generic Machine Learning Approach: A Danish National Health Register Study
Mie Sylow LILJENDAHL1, Kristina IBLER2, Christian VESTERGAARD3, Lone SKOV1,4, Pavika JAIN5, Jan Håkon RUDOLFSEN6, Ann HÆRSKJOLD2 and Mathias TORPET7
1Department of Dermatology and Allergy, Copenhagen University Hospital–Herlev and Gentofte, Copenhagen, 2Department of Dermatology, Bispebjerg Hospital, Copenhagen University Hospital, Copenhagen, 3Department of Dermatology, Aarhus University Hospital, Aarhus, 4Department of Clinical Medicine, Faculty of Health and Medical Sciences, University of Copenhagen, Copenhagen, 5Market Access, Sanofi Denmark A/S, 6EY, Frederiksberg, and 7Medical Affairs, Sanofi Denmark A/S, Denmark
Atopic dermatitis is a chronic skin disease, causing itching and recurrent eczematous lesions. In Danish national register data, adults with atopic dermatitis can only be identified if they have a hospital-diagnosed atopic dermatitis. The purpose of this study was to develop a machine learning model to identify all patients with atopic dermatitis by proxy, using data for contacts with primary care, prescription medication, and hospital contacts not related to skin diseases. Individuals redeeming a prescription for dermatological preparations were extracted as potential patients with atopic dermatitis. Individuals with a registered hospital diagnosis of atopic dermatitis were classified as “Known AD”, “Other skin disease” (registrations of other dermatological diagnosis codes indicating other skin disease), or “Uncertain AD status”’ (no hospital diagnosis registered). Patients categorized as “Known AD” and “Other skin disease” were used to develop the model. All uses of healthcare services 2 years prior to hospital diagnosis were used as potential predictors. The data were split into training and validation sets (70/30). From 1996 to 2022, 385,135 individuals had uncertain atopic dermatitis status. The most important predictors were corticosteroid prescriptions for dermatological use, consultations with dermatologist, and age. Of the 385,135 individuals, the model predicted that 230,522 individuals likely have atopic dermatitis.
SIGNIFICANCE
Atopic dermatitis is a chronic skin condition characterized by itching and recurring rashes. In Denmark, adults with atopic dermatitis are identified only if diagnosed in a hospital. This study developed a machine learning model to identify all atopic dermatitis patients using data from primary care visits, prescriptions, and unrelated hospital contacts. We analysed individuals who redeemed skin treatment prescriptions from 1996 to 2022. Key predictors included corticosteroid prescriptions, dermatologist consultations, and age. The model estimated that 230,522 individuals likely have atopic dermatitis. This study introduces the first machine learning model for identifying the adult atopic dermatitis population in Denmark, applicable to similar settings.
Key words: atopic dermatitis; machine learning; healthcare register; prediction.
Citation: Acta Derm Venereol 2025; 105: adv42250. DOI: https://doi.org/10.2340/actadv.v105.42250.
Copyright: © 2025 The Author(s). Published by MJS Publishing, on behalf of the Society for Publication of Acta Dermato-Venereologica. This is an Open Access article distributed under the terms of the Creative Commons Attribution-NonCommercial 4.0 International License (https://creativecommons.org/licenses/by-nc/4.0/).
Submitted: Oct 14, 2024. Accepted after revision: Apr 15, 2025. Published: May 13, 2025
Corr: Mie Sylow Liljendahl, Department of Dermatology and Allergy, Copenhagen University Hospital–Herlev and Gentofte, Denmark. E-mail: mie.sylow.liljendahl.01@regionh.dk
Competing interests and funding: MSL reports no conflict of interest. KI has been a paid speaker for Sanofi and Leo Pharma and has been consulted or served on Advisory Boards with Sanofi, Leo Pharma, Eli Lilly, and Almirall. She has served as an investigator for Sanofi, Leo Pharma, and Almirall and has received research and educational grants from Almirall, Region Zealand’s Research Fund, Birgit and Svend Igor Pock-Steens Fond, Kgl. Hofbundtmager Aage Bang Foundation, and the Danish Working Environment Research Fund. CV has been a paid consultant for Almirall, LEO Pharma, Sanofi Genzyme, Pfizer, and Abbvie. He has participated on a Data Safety Monitoring Board or Advisory Board for OM Pharma and served as the president of the Nordic Dermatology Association. LS has been a paid speaker for AbbVie, Eli Lilly, Novartis, Pfizer, Sanofi, and LEO Pharma, and has been a consultant or has served on Advisory Boards with AbbVie, Janssen, Novartis, Eli Lilly, Boehringer Ingelheim, LEO Pharma, UCB, Almirall, Bristol-Myers Squibb, and Sanofi. She has served as an investigator for AbbVie, Amgen, Sanofi, Janssen, Boehringer Ingelheim, Eli Lilly, Novartis, and LEO Pharma, and has received research and educational grants/funding from LEO Foundation, Kgl. Hofbundtmager Aage Bang Foundation, Novartis, Sanofi, Bristol-Myers Squibb, Janssen, Almirall, and LEO Pharma. PJ is a Sanofi employee and may hold shares and/or stock options in the company. JHR is an employee of EY. EY received funding from Sanofi to train and validate the machine learning model, as well as for medical writing assistance for this manuscript. AH has served on an Advisory board with Almirall. MT is a Sanofi employee and may hold shares and/or stock options in the company.
This study was funded by Sanofi, with medical writing assistance provided by Jens Olsen from EY.
INTRODUCTION
A topic dermatitis (AD) is a chronic inflammatory skin disease that causes recurring eczematous lesions and itching, which impair sleep, concentration, and quality of life (1–4). AD is caused by interactions between genetics, immunology, and environmental factors (5, 6). The prevalence of AD is uncertain but assumed to be 2–8% among European adults (7). The variation in the prevalence can be attributed to different diagnostic criteria and heterogeneous study designs (7). Moreover, identification of AD is challenging as it shares symptoms with many other skin diseases (8).
In Denmark, individuals with mild-to-moderate AD are treated in a primary healthcare setting and therefore not observable with diagnosis codes in the registers. Steroid-based treatments are used for many other similar skin conditions and are therefore unsuitable as an identifier for individuals with mild-to-moderate AD.
For moderate-to-severe AD, ICD-10 (International Statistical Classification of Diseases 10th Revision) diagnosis codes are registered for all inpatient and outpatient hospital contacts in Danish registers. Thus, these patients can be identified in the Danish National Patient Register (NPR) (9) by ICD-10 code L20 (including sublevels).
Using hospital contacts and the redemptions of AD-relieving medicines (by ATC codes), Henriksen et al. (10) developed an identification algorithm applicable to children with AD. Stensballe et al. (11) validated the algorithm, but Ortsäter et al. evaluated and concluded that the algorithm is not suited to identify adults with AD (12).
Even with access to nationwide healthcare registers, identification of patients with mild-to-moderate AD is challenging. Given the lack of algorithms identifying adults with mild-to-moderate AD, the aim of this study was to develop a machine learning model to identify adults with AD who are not identified in the NPR. The model is based on register data from primary and hospital healthcare contacts and the use of prescription medications. The generic machine learning framework can be applied to similar settings elsewhere or for conditions with similar properties.
Materials and methods
All Danish residents are assigned a unique personal identification ID at birth or upon immigration. This ID facilitates the secure linkage of data across national Danish registers. All individuals recorded as collecting a prescription for treatment of skin disorders between 1994 and 2021 in the Danish Register of Medicinal Product Statistics (13) were included. It was assumed that an individual with AD would necessarily have redeemed at least 1 relevant prescription (presented in Table SI).
Individuals recorded with an ICD-10 code L20 (including sublevels) in the NPR were classified as “Known AD”. If the individual was recorded with a diagnosis of other skin diseases in the NPR, they were classified as a “Other skin disease” (criteria presented in Table SII). Individuals not recorded with any dermatological hospital diagnosis were classified as “Uncertain AD status” (Fig. 1).
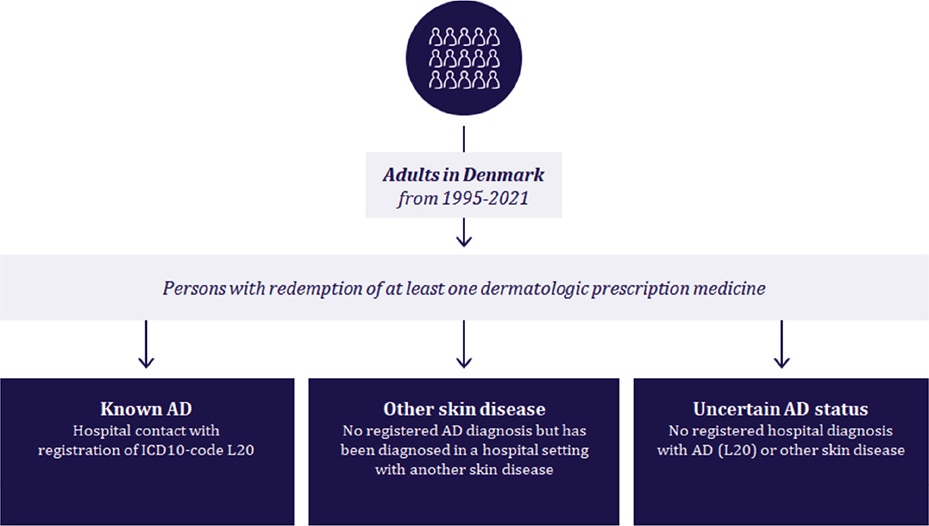
Fig. 1. Selection of individuals for the prediction model.
Age and region of residence were extracted from the Danish Central Person Register for all individuals. Services provided in the primary care sector were extracted from the Danish National Health Service Register (14). The population identified with “Other skin disease” was chosen as the control group. The aim of the model was to distinguish between “Known AD” and “Other skin disease”, implying that the “Uncertain AD status” population should be categorized as “Predicted AD” or “Predicted Other skin disease”.
Statistical methods
The model was trained to distinguish between “Known AD” and “Other skin disease”. The model was then applied to estimate the probability of an individual having AD in the “Uncertain AD status” population.
Identifying predictors. In training, all contacts with hospital care, primary care, and prescription medicine 2 years prior to dermatologically related hospital contact were included. The frequency of each service or redeemed prescription was aggregated for everyone during these 2 years, resulting in 8,990 variables.
For each of the count variables, a new dichotomized variable was made, reflecting each level of redeemed prescriptions or contacts. For example, if a prescription for drug A was collected 0–2 times, then this variable was split into 3 new variables (A0, A1, A2), indicating the number of collections an individual made in the period (one-hot encoding).
The training set contained 70% of the observations, while the remaining 30% of observations were used to assess the external validity of the model. The training and validation set were split at random. The data structure is illustrated in Fig. 2.
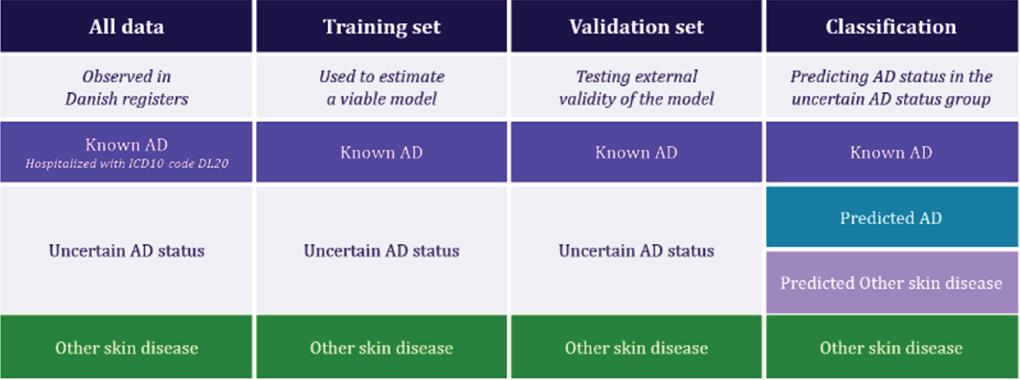

Fig. 2. Visualization of dataset structure and overview of the analytic approach (moving from a sub-sample of individuals with “Uncertain AD status” to a classification of all individuals). “Known AD”: hospital contact with registration of ICD10-code L20; “Other skin disease”: no registered AD diagnosis but registration of another skin disease diagnosis in a hospital setting; “Uncertain AD status”: redemption of at least 1 dermatologic prescription medicine but no registered hospital diagnosis with AD (L20) or other skin disease.
The first step in identifying suitable predictors was to remove features with low variation. Variables where features had the same value more than 90% of the time were removed. The second step in variable reduction was to include only features with a correlation to the outcome. Features with a p-value ≤ 0.10 in a univariate linear regression model were retained for further model development.
Finally, we applied recursive feature elimination, where the variable with lowest predictive power was identified and removed. After removal, the model was re-estimated, and the process was repeated until only 1 predictor remained.
The predictive power of the variables was evaluated by mean decrease in accuracy in a random forest (RF) model (15). To estimate one RF model, we applied 500 iterations (see Fig. 3). Overall accuracy (the percentage of correct predictions) and area under curve (AUC) were stored for each iteration.
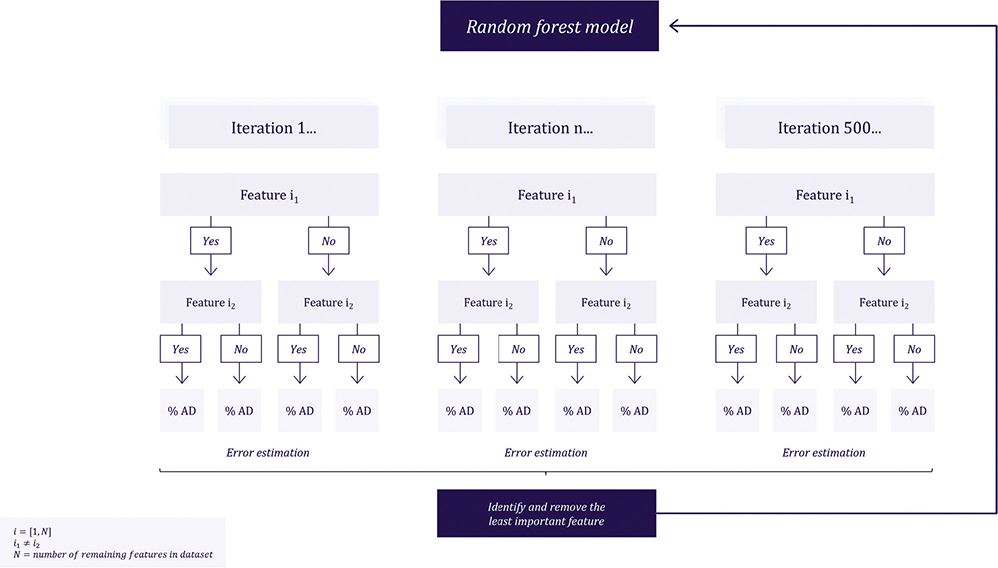
Fig. 3. Illustration of random forest model process for feature elimination involving 500 iterations.
Final model. The RF models with fewer than 25 features were evaluated further. Expert opinion was used to omit features with no clinical foundation and to include clinically meaningful variables with a clear indication of AD that were not included after data-driven assessment.
An evaluation of the skewed distribution of “Known AD” and “Other skin disease” was conducted, as 83% of observations were classified as “Known AD”. Extrapolating this distribution to the observations in “Uncertain AD status” would result in an unrealistic high prevalence of AD in the Danish population. Therefore, class-based weighting was assigned such that the sum of weights for the “Known AD” population and “Other skin disease” population totalled 1. The final model was estimated using an RF model in the ranger package (version 0.15.1) in R (version 4.3.2) (R Foundation for Statistical Computing, Vienna, Austria). External validity was evaluated in the validation set based on overall accuracy, positive and negative prediction value, calibration slope, and AUC. Confidence intervals for overall accuracy and AUC were obtained by bootstrapping, with 1,000 repetitions of 250 random draws.
Validation of “Uncertain AD status”. It was not possible to conclusively validate the accuracy of the prediction among individuals with “Uncertain AD status” in the final model. Hence, 2 alternative strategies were applied to inform on the validity of the predictions. First, a comparison of predicted AD characteristics with previously published studies on mild to severe AD was conducted. Second, a considerable number of individuals became adults during the observation period. For these individuals, we estimated how many were identified as having AD as children in the validated Henriksen algorithm and were also identified by our machine learning model after they became adults.
Results
The process of identifying the populations with “Known AD”, “Other skin disease”, and “Uncertain AD status” is presented in Fig. 1, with summary statistics presented in Table I. Summary statistics for the “Known AD” and “Other skin disease” populations were calculated at the time of first hospital contact. Summary statistics for the “Uncertain AD status” population were calculated in the individuals’ last year of observation.
The “Known AD” population was somewhat younger than the “Other skin disease” population. Moreover, a higher proportion of the “Known AD” population was identified early in the study period compared with those with “Other skin disease”. We observed that more women than men were diagnosed with a dermatological condition in a hospital setting. The sex distribution was less skewed in the “Uncertain AD status” population.
Model input
The data-driven modelling process identified 16 variables relevant for prediction. Eight of these were omitted due to their theoretically spurious relation to AD diagnosis. Furthermore, use of “tacrolimus” and “pimecrolimus” had been excluded from the data-driven model approach due to low variation. However, as these are used uniquely for individuals with AD in Denmark they were reintroduced to the model.
By combining data-driven insights with expert knowledge, we identified 10 variables suitable for distinguishing between “Known AD” and “Other skin disease”. The included and excluded variables are presented in Table II.
Model accuracy and validation
Model metrics are presented in Table III. The training model achieved an overall accuracy of 85% (95% confidence interval [CI]: 85–86%]. The accuracy in the validation set was 72% (CI: 70–73%), with AUC = 0.71 (CI: 0.66–0.76) and a calibration slope of 1.09, indicating a slight under-fitting. This suggests that the model underestimates the risk of higher probabilities and overestimates the risk of lower probabilities. Both the training and validation sets demonstrated balanced predictions, based on positive and negative predicted value.
Of 385,135 individuals with “Uncertain AD status”, the model predicted that 230,522 individuals (59.9%) had AD. Including the individuals diagnosed with AD in a hospital setting (“Known AD”), we identified 243,500 adult individuals with AD during the study period. Summary statistics on predicted AD and predicted “Other skin disease” are provided in Table IV. A relatively large number of individuals with AD were identified early in the study period, as these individuals were likely prevalent at the beginning of the study period (1994) and should therefore be included from the outset.
Of the 243,500 adult individuals identified in this study, 191,394 individuals were alive and residing in Denmark by the end of 2021. This corresponds to a prediction of adults with AD in Denmark of 4,054 per 100,000.
To validate the accuracy of our model, we considered the population transitioning from children to adults during the study period and compared it with the verified Henriksen et al. algorithm. Of 385,175 individuals with “uncertain AD status”, 163,403 children became adults in the study period who were previously identified by the Henriksen et al. algorithm. The machine learning model developed in this study identified 160,967 of these individuals as “AD” individuals in adulthood.
DISCUSSION
The burden of AD in Denmark is unknown. In this study, we have proposed a method to identify individuals with AD who have not been diagnosed in hospital care, based on patterns of health service consumption observed in national health registers. The model achieved an accuracy of 85% (validation: 72%) and was able to identify individuals on a comparable basis to related studies (10). The accuracy fit was within the range considered acceptable for application in clinical care (16). In the validation set, the model outperformed the Henriksen et al. (10) algorithm for identifying adults with AD (12), even when the -Henriksen et al. algorithm is adapted to this purpose.
The findings of Ortsäter et al. (12) indicate that retro-diagnostic algorithms must be adapted to the healthcare system in which they are applied. We do not expect our model to achieve acceptable accuracy if applied in populations outside of Denmark. However, the methodology described could be transferred to other healthcare systems with similar results.
The performance metrics suggest underfitting, indicated by the calibration slope, which implies that the model underestimates risk for higher probabilities and overestimates risk for lower probabilities. This makes the model less reliable in predicting extreme values, although it remains reasonably effective at identifying true negatives. Meanwhile, the difference in accuracy between the training set and the validation set indicates overfitting. However, the balanced validation demonstrated high accuracy with a narrow confidence interval, indicating that the model maintains acceptable performance across both classes, despite the slight miscalibration. This suggests that the model is still a reliable tool for classification. A balance between these 2 contradictory effects was attempted by reducing the number of predictors, based on both data-driven methods and expert opinion.
The topic of overfitting and underfitting should be considered in the context of the extent to which AD treatment patterns can be observed in register data, and how accurate a perfect model can be, given the available data. AD exhibits traits of preference-sensitive care (17), i.e., conditions where treatment is subject to physicians’ perception and patients’ preference. If that is the case, any model will suffer similar shortcomings, highlighting the need for a methodological framework to identify AD cases in each healthcare system, rather than standardized models.
We recognize the potential for misreporting of information in the registers, and the authors did not have the opportunity to control for this. The Danish national registers are considered to be of high quality and are frequently used in scientific studies. Drug adherence could not be investigated.
The analysis is based on a comprehensive dataset containing almost 9,000 features. With the aim of creating a functional model with external validity, it was necessary to reduce the input to develop a more applicable model. By initially applying a data-driven approach, the researcher’s degrees of freedom were minimized. However, this approach will ignore any real-world expertise, which is why expert evaluation from practising physicians was incorporated into the model-building process.
While this process ensures a robust variable selection with a balanced approach to model specification, multiple aspects will influence the overall accuracy. For example, a forward-stepping approach using AIC (Akaike Information Criterion) or BIC (Bayesian Information Criterion) as metrics for variable selection could have affected the choice of input factors for the final model. Moreover, other variable transformations than one-hot encoding could have been used. The one-hot encoding method removes any ordinal information in variable selection, which may have resulted in the omission of important information. However, the bias in the method towards high cardinality might have been advantageous in this context, as the model was targeted at a large general skin disease population, influencing the evaluations of the input factors. Lastly, the choice of using RF to estimate the final model may have impacted the accuracy of predictions.
There is the potential for some individuals with psoriasis to be classified as having “Known AD”. In total, 583 individuals (0.2%) out of the 385,135 with “uncertain AD status” collected at least 1 prescription medication with the ATC code D05 (antipsoriatics, including subgroups). Of the 583 individuals, 422 were predicted as “AD”. However, 71.3% of these redeemed only 1 prescription for antipsoriatic medication. We do not consider it to have influenced the results.
The advantage of the machine learning method demonstrated in this study is that large datasets can be handled efficiently, allowing for the creation of data-driven models with sufficient accuracy to be applied in an epidemiological setting, and flexible enough to be adapted to a new context. For example, the variable transformation might not be suited to another context, and the recursive feature elimination could be replaced with forward-stepping model building, and the final RF model could be replaced by alternative models.
For the current case, the variable transformation and subsequent feature elimination was deemed suited due to the wide initial dataset, and not to overlook potential predictors due to researchers’ degrees of freedom. Moreover, the application of the RF model implicitly accounts for interaction effects, which is important when evaluating the potentially complex treatment patterns in a wide population.
In conclusion, this study presents the first machine learning model to identify the adult AD population in Denmark. The model was developed and validated specifically for the healthcare context of Denmark and verified by clinical experts. The method applied can be implemented in other countries with similar national registers data. Unlike conventional register data processing, this machine learning model can identify complex patterns and interactions in national registers, enabling a more comprehensive and precise identification of AD cases. Identifying all individuals with AD outside the hospital sector enables potential research on the entire AD population, including regional differences in healthcare utilization and potential identification of health inequalities.
ACKNOWLEDGEMENTS
IRB approval status
The data that support the findings of this study are available from Statistics Denmark’s Research Service. The study was complied with the regulations and instructions set up by Statistics Denmark. We used only anonymized data, and present data only in aggregate and anonymous form. We did not contact or require any active participation from study participants. Ethics committee approval and written informed consent are not required for register-based research, according to Danish law.
REFERENCES
- Silverberg JI, Gelfand JM, Margolis DJ, Boguniewicz M, Fonacier L, Grayson MH, et al. Pain is a common and burdensome symptom of atopic dermatitis in United States adults. J Allergy Clin Immunol Pract 2019; 7: 2699–2706. https://doi.org/10.1016/j.jaip.2019.05.055
- Yu SH, Attarian H, Zee P, Silverberg JI. Burden of sleep and fatigue in US adults with atopic dermatitis. Dermatitis 2016; 27: 50–58. https://doi.org/10.1097/DER.0000000000000161
- Blome C, Radtke MA, Eissing L, Augustin M. Quality of life in patients with atopic dermatitis: disease burden, measurement, and treatment benefit. Am J Clin Dermatol 2016; 17: 163–169. https://doi.org/10.1007/s40257-015-0171-3
- Ali F, Vyas J, Finlay A. Counting the burden: atopic dermatitis and health-related quality of life. Acta Derm Venereol 2020; 100: 5766. https://doi.org/10.2340/00015555-3511
- Malik K, Heitmiller KD, Czarnowicki T. An update on the pathophysiology of atopic dermatitis. Dermatol Clin 2017; 35: 317–326. https://doi.org/10.1016/j.det.2017.02.006
- Bieber T, Paller AS, Kabashima K, Feely M, Rueda MJ, Ross Terres JA, et al. Atopic dermatitis: pathomechanisms and lessons learned from novel systemic therapeutic options. J Eur Acad Dermatol Venereol 2022; 36:1432–1449. https://doi.org/10.1111/jdv.18225
- Bylund S, Kobyletzki L, Svalstedt M, Svensson Å. Prevalence and incidence of atopic dermatitis: a systematic review. Acta Derm Venereol 2020; 100: 320–329. https://doi.org/10.2340/00015555-3510
- Barrett M, Luu M. Differential diagnosis of atopic dermatitis. Immunol Allergy Clin North Am 2017; 37: 11–34. https://doi.org/10.1016/j.iac.2016.08.009
- Lynge E, Sandegaard JL, Rebolj M. The Danish National Patient Register. Scand J Public Health 2011; 39: 30–33. https://doi.org/10.1177/1403494811401482
- Henriksen L, Simonsen J, Haerskjold A, Linder M, Kieler H, Thomsen SF, et al. Incidence rates of atopic dermatitis, asthma, and allergic rhinoconjunctivitis in Danish and Swedish children. J Allergy Clin Immunol 2015; 136: 360–366. https://doi.org/10.1016/j.jaci.2015.02.003
- Stensballe LG, Klansø L, Jensen A, Haerskjold A, Thomsen SF, Simonsen J. The validity of register data to identify children with atopic dermatitis, asthma or allergic rhinoconjunctivitis. Pediatr Allergy Immunol 2017; 28: 535–542. https://doi.org/10.1111/pai.12743
- Ortsäter G, De Geer A, Geale K, Rieem Dun A, Lindberg I, Thyssen JP, et al. Validation of patient identification algorithms for atopic dermatitis using healthcare databases. Dermatol Ther (Heidelb) 2022; 12: 545–559. https://doi.org/10.1007/s13555-021-00670-1
- Wallach Kildemoes H, Toft Sørensen H, Hallas J. The Danish National Prescription Registry. Scand J Public Health 2011; 39: 38–41. https://doi.org/10.1177/1403494810394717
- Sahl Andersen J, De Fine Olivarius N, Krasnik A. The Danish National Health Service Register. Scand J Public Health 2011; 39: 34–37. https://doi.org/10.1177/1403494810394718
- Breiman L. Random forests. Machine Learning 2001; 45: 5–32. https://doi.org/10.1023/A:1010933404324
- Alba AC, Agoritsas T, Walsh M, Hanna S, Iorio A, Devereaux PJ, et al. Discrimination and calibration of clinical prediction models: users’ guides to the medical literature. JAMA 2017; 318: 1377–1384. https://doi.org/10.1001/jama.2017.12126
- Wennberg JE. Unwarranted variations in healthcare delivery: implications for academic medical centres. BMJ 2002; 325: 961–964. https://doi.org/10.1136/bmj.325.7370.961